🧬Genetic Material: DNA, RNA, Gene Concepts, and Genetic Code
Master DNA and RNA as genetic material, gene concepts (operon, cistron), and the genetic code — with agricultural examples, comparison tables, and exam-focused mnemonics.
Why Genetic Material Matters in Agriculture
When scientists developed Bt cotton by inserting a bacterial gene into cotton DNA, or when breeders use molecular markers to select disease-resistant rice lines without waiting for the disease to appear, they are working directly with genetic material. The discovery that DNA (not protein) carries hereditary information transformed agriculture — enabling marker-assisted selection, genetic engineering, and genomics-based crop improvement. Understanding DNA, RNA, genes, and the genetic code is essential for modern plant breeding and biotechnology.
What Is Genetic Material?
Genetic material consists of nucleic acids — the molecules that store and transmit hereditary information.
| Nucleic Acid | Full Name | Role |
|---|---|---|
| DNA | Deoxyribose Nucleic Acid | Primary genetic material in all cellular organisms |
| RNA | Ribose Nucleic Acid | Genetic material in some viruses; carries DNA’s instructions in all organisms |
Key Discoveries
| Scientist(s) | Year | Discovery |
|---|---|---|
| Miescher | 1868 | First isolated nucleic acids (“nuclein”) from WBC pus |
| Avery, MacLeod & McCarty | 1944 | Proved DNA (not protein) is the genetic material (transforming principle in E. coli) |
| A. Kornberg | 1959 | First in vitro synthesis of DNA (discovered DNA polymerase; Nobel Prize) |
| S. Ochoa | 1959 | In vitro synthesis of RNA (discovered polynucleotide phosphorylase; Nobel Prize) |
| H.G. Khorana & K.L. Agrawal | — | First artificial synthesis of a complete functional gene (alanine tRNA gene from yeast) |
Exam tip: The Avery-MacLeod-McCarty experiment (1944) is one of the most frequently asked discoveries. Remember: “AMM proved DNA, not protein”.
Gene Concepts — Evolution of Understanding
One Gene–One Enzyme Hypothesis
- Proposed by
Beadle & Tatum(1943) using biochemical mutants of Neurospora crassa (red bread mould). - Each gene codes for one specific enzyme.
- Later refined to “one gene–one polypeptide” (not all gene products are enzymes; some proteins have multiple polypeptide chains coded by different genes).
Progression of the concept:
- One gene → One enzyme
- One gene → One protein
- One gene → One polypeptide chain
- One cistron → One polypeptide
Operon Concept
- Given by
Jacob & Monod(Nobel Prize 1965). - Explains how gene expression is regulated in prokaryotes.
- Classic example: lac operon in E. coli — genes are switched on/off together in response to environmental signals (e.g., presence of lactose).
Gene Fine Structure
- Established by
Benzerusing bacteriophage T4. - Showed that genes are not indivisible — they can be mapped into smaller functional regions.
Benzer’s Three Units of the Gene
| Unit | Definition | Size | Key Fact |
|---|---|---|---|
Recon | Smallest unit of recombination | 1–2 nucleotide pairs | Recombination can occur between adjacent nucleotides |
Muton | Smallest unit of mutation | Single nucleotide pair (smallest) | Even a point mutation can alter gene function |
Cistron | Functional unit of the gene | Hundreds of nucleotide pairs (largest) | Equivalent to “gene” in modern usage; codes for one polypeptide |
Mnemonic: Size order = Recon < Muton < Cistron (alphabetical order R-M-C matches smallest to largest).
Structure of DNA
Watson-Crick Model (1953)
- Proposed by J.D. Watson & F.H.C. Crick.
- X-ray diffraction data by Wilkins and Rosalind Franklin (Photo 51).
- Nobel Prize (1962): Watson, Crick, and Wilkins.
Double Helix Features
| Parameter | Value |
|---|---|
| Two antiparallel strands | 5’→3’ and 3’→5’ |
| Backbone | Sugar-phosphate on the outside |
| Bases | On the inside; form hydrogen bonds |
| A–T | 2 hydrogen bonds |
| G–C | 3 hydrogen bonds (more thermally stable) |
| Distance between base pairs | 3.4 Å |
| Base pairs per turn | 10 |
| Length per turn | 34 Å |
| Helix diameter | 20 Å |

Chargaff’s Rules
- A = T and G = C; total purines (A+G) = total pyrimidines (T+C).
- (A+T)/(G+C) =
Base pair ratio— unique to each species (biochemical fingerprint). - The two strands are complementary (not identical) and run antiparallel.
- Knowing one strand’s sequence automatically reveals the other — this is the basis of DNA replication and transcription.
Agricultural application: Base pair ratio and G-C content affect DNA melting temperature (Tm), which is critical when designing PCR primers for molecular markers used in crop breeding (SSR, RAPD, SNP markers).
Structure of RNA
| Feature | DNA | RNA |
|---|---|---|
| Sugar | Deoxyribose | Ribose |
| Strands | Double-stranded (helical) | Usually single-stranded (can fold into 3D shapes) |
| Bases | A, T, G, C | A, U, G, C |
| Genetic role | Primary genetic material (all cellular organisms) | Genetic material in some plant viruses and bacteriophages |
| Virus | Type of RNA |
|---|---|
| Plant Viruses | |
| Turnip yellow mosaic virus (TYMC) | Single stranded |
| Wound tumour | Double stranded |
| Animal viruses | |
| Influenza virus | Single stranded |
| Rous Sarcoma | Single stranded |
| Poliomyelitis | Single stranded |
| Reovirus | Double stranded |
| Bacteriophages | |
| MS 2, F 2, r 17 | Single stranded |

Types of RNA
| Type | Category | Feature |
|---|---|---|
| Genetic RNA | Viral genomes | Self-replicating via RNA-dependent RNA synthesis (enzyme: RdRp) |
| Non-genetic RNA | mRNA, tRNA, rRNA | Synthesised on DNA template; carry out DNA’s instructions |
- In organisms that have both DNA and RNA, the RNA performs non-genetic roles (messenger, transfer, ribosomal).
- tRNA is double-stranded but non-helical (clover-leaf model).
Agricultural example: Many devastating crop diseases are caused by RNA viruses — rice tungro, tomato spotted wilt, wheat streak mosaic. Understanding RNA replication (RdRp) is the basis for developing antiviral strategies in crops.
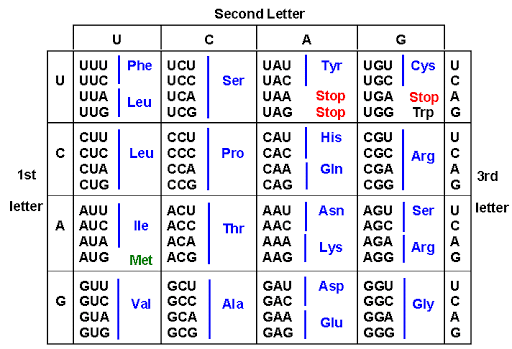
The Genetic Code
Deciphering the Code
- Holley, Khorana, and Nirenberg — Nobel Prize in Physiology or Medicine (1968).
- DNA’s genetic information is written in a language of 4 bases (A, T, G, C) but must code for 20 amino acids in proteins.
- The coded message on DNA is called a cryptogram; it is transmitted via mRNA to ribosomes.
Why Triplet Code?
| Code Type | Combinations | Sufficient for 20 amino acids? |
|---|---|---|
| Singlet (1 base) | 4¹ = 4 | No |
| Doublet (2 bases) | 4² = 16 | No |
| Triplet (3 bases) | 4³ = 64 | Yes (with 44 redundant codons) |
An anticodon is the complementary triplet on tRNA that matches the mRNA codon, ensuring the correct amino acid is delivered.

Five Properties of the Genetic Code
IMPORTANT
Memorise these five properties — they are tested repeatedly in competitive exams.
| Property | Meaning | Example |
|---|---|---|
| Triplet | Each codon = 3 bases | Minimum requirement to code 20 amino acids |
| Degenerate | Multiple codons for same amino acid (synonyms) | Arg, Ser, Leu each have 6 codons; all CC- codons = proline |
| Non-overlapping | Each base belongs to only one codon | No base is shared between adjacent codons |
| Comma-less | No spacers or punctuation between codons | Ribosome reads continuously, 3 bases at a time |
| Universal | Same code in all organisms | From bacteria to humans; minor exceptions in mitochondrial DNA |
Additional property:
- Ambiguous — under abnormal conditions (e.g., streptomycin), a codon may code for a different amino acid. This is NOT a normal feature of translation.
Mnemonic for code properties: “TDN-CU” — Triplet, Degenerate, Non-overlapping, Comma-less, Universal.
Start and Stop Codons
| Type | Codons | Name | Function |
|---|---|---|---|
| Start | AUG | — | Initiates translation; codes for methionine |
| Stop | UAA | Ochre | Terminates translation |
| Stop | UAG | Amber | Terminates translation |
| Stop | UGA | Opal | Terminates translation |
Mnemonic for stop codons: “U Are Annoying, U Are Gone, U Go Away” — UAA, UAG, UGA.
Degeneracy benefit in agriculture: Because of code degeneracy, many point mutations in the third codon position are silent (do not change the amino acid). This provides a buffer against harmful mutations in crop genomes, contributing to genetic stability across generations.
Explore More
- Genetic Code and Codons Explained
- Transcription and Translation Overview
- Gene Expression: From DNA to Protein
Summary Table
| Topic | Key Fact | Exam Pointer |
|---|---|---|
| DNA as genetic material | Proved by Avery, MacLeod & McCarty (1944) | Transforming principle in E. coli |
| Nucleic acids first isolated | Miescher (1868) from WBC pus | Called it “nuclein” |
| In vitro DNA synthesis | A. Kornberg (Nobel 1959) | Discovered DNA polymerase |
| In vitro RNA synthesis | S. Ochoa (Nobel 1959) | Polynucleotide phosphorylase |
| Artificial gene synthesis | Khorana & Agrawal | Alanine tRNA gene from yeast |
| One gene–one enzyme | Beadle & Tatum (1943) on Neurospora | Refined to one gene–one polypeptide |
| Operon concept | Jacob & Monod (Nobel 1965) | lac operon in E. coli |
| Gene fine structure | Benzer (phage T4) | Recon < Muton < Cistron |
| DNA model | Watson & Crick (1953; Nobel 1962) | Double helix; 3.4 Å/bp; 10 bp/turn |
| Chargaff’s Rules | A=T, G=C | Base pair ratio is species-specific |
| Genetic code | Triplet, degenerate, universal | 64 codons for 20 amino acids |
| Start codon | AUG (methionine) | Initiates translation |
| Stop codons | UAA, UAG, UGA | Ochre, Amber, Opal |
| Degeneracy | Multiple codons per amino acid | Wobble position provides mutation buffer |
Summary Cheat Sheet
| Concept / Topic | Key Details |
|---|---|
| Nucleic acids first isolated by | Miescher (1868) — “nuclein” from WBC pus |
| DNA proved as genetic material | Avery, MacLeod & McCarty (1944) |
| In vitro DNA synthesis | A. Kornberg (Nobel 1959); discovered DNA polymerase |
| In vitro RNA synthesis | S. Ochoa (Nobel 1959); polynucleotide phosphorylase |
| Artificial gene synthesis | Khorana & Agrawal — alanine tRNA gene from yeast |
| One gene–one enzyme | Beadle & Tatum (1943) on Neurospora crassa |
| Modern refinement | One gene → one polypeptide |
| Operon concept | Jacob & Monod (Nobel 1965); lac operon in E. coli |
| Benzer’s units (size order) | Recon (1–2 bp) < Muton (1 bp) < Cistron (hundreds bp) |
| Recon | Smallest unit of recombination |
| Muton | Smallest unit of mutation |
| Cistron | Functional unit = gene; codes for one polypeptide |
| DNA double helix | Watson & Crick (1953); Nobel Prize 1962 |
| X-ray data by | Wilkins & Rosalind Franklin (Photo 51) |
| A–T = 2 H-bonds | G–C = 3 H-bonds (more stable) |
| DNA dimensions | 3.4 Å/bp, 10 bp/turn, 34 Å/turn, 20 Å diameter |
| Chargaff’s Rules | A=T, G=C; (A+T)/(G+C) = species-specific ratio |
| RNA vs DNA | RNA: ribose, single-stranded, uracil replaces thymine |
| RdRp | RNA-dependent RNA polymerase; replicates viral RNA genomes |
| Genetic code deciphered by | Holley, Khorana, Nirenberg (Nobel 1968) |
| Triplet code | 4³ = 64 codons for 20 amino acids |
| Code properties | Triplet, Degenerate, Non-overlapping, Comma-less, Universal |
| Start codon | AUG (methionine) |
| Stop codons | UAA (ochre), UAG (amber), UGA (opal) |
Pro Content Locked
Upgrade to Pro to access this lesson and all other premium content.
₹2388 billed yearly
- All Agriculture & Banking Courses
- AI Lesson Questions (100/day)
- AI Doubt Solver (50/day)
- Glows & Grows Feedback (30/day)
- AI Section Quiz (20/day)
- 22-Language Translation (30/day)
- Recall Questions (20/day)
- AI Quiz (15/day)
- AI Quiz Paper Analysis
- AI Step-by-Step Explanations
- Spaced Repetition Recall (FSRS)
- AI Tutor
- Immersive Text Questions
- Audio Lessons — Hindi & English
- Mock Tests & Previous Year Papers
- Summary & Mind Maps
- XP, Levels, Leaderboard & Badges
- Generate New Classrooms
- Voice AI Teacher (AgriDots Live)
- AI Revision Assistant
- Knowledge Gap Analysis
- Interactive Revision (LangGraph)
🔒 Secure via Razorpay · Cancel anytime · No hidden fees
Why Genetic Material Matters in Agriculture
When scientists developed Bt cotton by inserting a bacterial gene into cotton DNA, or when breeders use molecular markers to select disease-resistant rice lines without waiting for the disease to appear, they are working directly with genetic material. The discovery that DNA (not protein) carries hereditary information transformed agriculture — enabling marker-assisted selection, genetic engineering, and genomics-based crop improvement. Understanding DNA, RNA, genes, and the genetic code is essential for modern plant breeding and biotechnology.
What Is Genetic Material?
Genetic material consists of nucleic acids — the molecules that store and transmit hereditary information.
| Nucleic Acid | Full Name | Role |
|---|---|---|
| DNA | Deoxyribose Nucleic Acid | Primary genetic material in all cellular organisms |
| RNA | Ribose Nucleic Acid | Genetic material in some viruses; carries DNA’s instructions in all organisms |
Key Discoveries
| Scientist(s) | Year | Discovery |
|---|---|---|
| Miescher | 1868 | First isolated nucleic acids (“nuclein”) from WBC pus |
| Avery, MacLeod & McCarty | 1944 | Proved DNA (not protein) is the genetic material (transforming principle in E. coli) |
| A. Kornberg | 1959 | First in vitro synthesis of DNA (discovered DNA polymerase; Nobel Prize) |
| S. Ochoa | 1959 | In vitro synthesis of RNA (discovered polynucleotide phosphorylase; Nobel Prize) |
| H.G. Khorana & K.L. Agrawal | — | First artificial synthesis of a complete functional gene (alanine tRNA gene from yeast) |
Exam tip: The Avery-MacLeod-McCarty experiment (1944) is one of the most frequently asked discoveries. Remember: “AMM proved DNA, not protein”.
Gene Concepts — Evolution of Understanding
One Gene–One Enzyme Hypothesis
- Proposed by
Beadle & Tatum(1943) using biochemical mutants of Neurospora crassa (red bread mould). - Each gene codes for one specific enzyme.
- Later refined to “one gene–one polypeptide” (not all gene products are enzymes; some proteins have multiple polypeptide chains coded by different genes).
Progression of the concept:
- One gene → One enzyme
- One gene → One protein
- One gene → One polypeptide chain
- One cistron → One polypeptide
Operon Concept
- Given by
Jacob & Monod(Nobel Prize 1965). - Explains how gene expression is regulated in prokaryotes.
- Classic example: lac operon in E. coli — genes are switched on/off together in response to environmental signals (e.g., presence of lactose).
Gene Fine Structure
- Established by
Benzerusing bacteriophage T4. - Showed that genes are not indivisible — they can be mapped into smaller functional regions.
Benzer’s Three Units of the Gene
| Unit | Definition | Size | Key Fact |
|---|---|---|---|
Recon | Smallest unit of recombination | 1–2 nucleotide pairs | Recombination can occur between adjacent nucleotides |
Muton | Smallest unit of mutation | Single nucleotide pair (smallest) | Even a point mutation can alter gene function |
Cistron | Functional unit of the gene | Hundreds of nucleotide pairs (largest) | Equivalent to “gene” in modern usage; codes for one polypeptide |
Mnemonic: Size order = Recon < Muton < Cistron (alphabetical order R-M-C matches smallest to largest).
Structure of DNA
Watson-Crick Model (1953)
- Proposed by J.D. Watson & F.H.C. Crick.
- X-ray diffraction data by Wilkins and Rosalind Franklin (Photo 51).
- Nobel Prize (1962): Watson, Crick, and Wilkins.
Double Helix Features
| Parameter | Value |
|---|---|
| Two antiparallel strands | 5’→3’ and 3’→5’ |
| Backbone | Sugar-phosphate on the outside |
| Bases | On the inside; form hydrogen bonds |
| A–T | 2 hydrogen bonds |
| G–C | 3 hydrogen bonds (more thermally stable) |
| Distance between base pairs | 3.4 Å |
| Base pairs per turn | 10 |
| Length per turn | 34 Å |
| Helix diameter | 20 Å |

Chargaff’s Rules
- A = T and G = C; total purines (A+G) = total pyrimidines (T+C).
- (A+T)/(G+C) =
Base pair ratio— unique to each species (biochemical fingerprint). - The two strands are complementary (not identical) and run antiparallel.
- Knowing one strand’s sequence automatically reveals the other — this is the basis of DNA replication and transcription.
Agricultural application: Base pair ratio and G-C content affect DNA melting temperature (Tm), which is critical when designing PCR primers for molecular markers used in crop breeding (SSR, RAPD, SNP markers).
Structure of RNA
| Feature | DNA | RNA |
|---|---|---|
| Sugar | Deoxyribose | Ribose |
| Strands | Double-stranded (helical) | Usually single-stranded (can fold into 3D shapes) |
| Bases | A, T, G, C | A, U, G, C |
| Genetic role | Primary genetic material (all cellular organisms) | Genetic material in some plant viruses and bacteriophages |
| Virus | Type of RNA |
|---|---|
| Plant Viruses | |
| Turnip yellow mosaic virus (TYMC) | Single stranded |
| Wound tumour | Double stranded |
| Animal viruses | |
| Influenza virus | Single stranded |
| Rous Sarcoma | Single stranded |
| Poliomyelitis | Single stranded |
| Reovirus | Double stranded |
| Bacteriophages | |
| MS 2, F 2, r 17 | Single stranded |

Types of RNA
| Type | Category | Feature |
|---|---|---|
| Genetic RNA | Viral genomes | Self-replicating via RNA-dependent RNA synthesis (enzyme: RdRp) |
| Non-genetic RNA | mRNA, tRNA, rRNA | Synthesised on DNA template; carry out DNA’s instructions |
- In organisms that have both DNA and RNA, the RNA performs non-genetic roles (messenger, transfer, ribosomal).
- tRNA is double-stranded but non-helical (clover-leaf model).
Agricultural example: Many devastating crop diseases are caused by RNA viruses — rice tungro, tomato spotted wilt, wheat streak mosaic. Understanding RNA replication (RdRp) is the basis for developing antiviral strategies in crops.
The Genetic Code
Deciphering the Code
- Holley, Khorana, and Nirenberg — Nobel Prize in Physiology or Medicine (1968).
- DNA’s genetic information is written in a language of 4 bases (A, T, G, C) but must code for 20 amino acids in proteins.
- The coded message on DNA is called a cryptogram; it is transmitted via mRNA to ribosomes.
Why Triplet Code?
| Code Type | Combinations | Sufficient for 20 amino acids? |
|---|---|---|
| Singlet (1 base) | 4¹ = 4 | No |
| Doublet (2 bases) | 4² = 16 | No |
| Triplet (3 bases) | 4³ = 64 | Yes (with 44 redundant codons) |
An anticodon is the complementary triplet on tRNA that matches the mRNA codon, ensuring the correct amino acid is delivered.

Five Properties of the Genetic Code
IMPORTANT
Memorise these five properties — they are tested repeatedly in competitive exams.
| Property | Meaning | Example |
|---|---|---|
| Triplet | Each codon = 3 bases | Minimum requirement to code 20 amino acids |
| Degenerate | Multiple codons for same amino acid (synonyms) | Arg, Ser, Leu each have 6 codons; all CC- codons = proline |
| Non-overlapping | Each base belongs to only one codon | No base is shared between adjacent codons |
| Comma-less | No spacers or punctuation between codons | Ribosome reads continuously, 3 bases at a time |
| Universal | Same code in all organisms | From bacteria to humans; minor exceptions in mitochondrial DNA |

Additional property:
- Ambiguous — under abnormal conditions (e.g., streptomycin), a codon may code for a different amino acid. This is NOT a normal feature of translation.
Mnemonic for code properties: “TDN-CU” — Triplet, Degenerate, Non-overlapping, Comma-less, Universal.
Start and Stop Codons
| Type | Codons | Name | Function |
|---|---|---|---|
| Start | AUG | — | Initiates translation; codes for methionine |
| Stop | UAA | Ochre | Terminates translation |
| Stop | UAG | Amber | Terminates translation |
| Stop | UGA | Opal | Terminates translation |
Mnemonic for stop codons: “U Are Annoying, U Are Gone, U Go Away” — UAA, UAG, UGA.
Degeneracy benefit in agriculture: Because of code degeneracy, many point mutations in the third codon position are silent (do not change the amino acid). This provides a buffer against harmful mutations in crop genomes, contributing to genetic stability across generations.
Explore More
- Genetic Code and Codons Explained
- Transcription and Translation Overview
- Gene Expression: From DNA to Protein
Summary Table
| Topic | Key Fact | Exam Pointer |
|---|---|---|
| DNA as genetic material | Proved by Avery, MacLeod & McCarty (1944) | Transforming principle in E. coli |
| Nucleic acids first isolated | Miescher (1868) from WBC pus | Called it “nuclein” |
| In vitro DNA synthesis | A. Kornberg (Nobel 1959) | Discovered DNA polymerase |
| In vitro RNA synthesis | S. Ochoa (Nobel 1959) | Polynucleotide phosphorylase |
| Artificial gene synthesis | Khorana & Agrawal | Alanine tRNA gene from yeast |
| One gene–one enzyme | Beadle & Tatum (1943) on Neurospora | Refined to one gene–one polypeptide |
| Operon concept | Jacob & Monod (Nobel 1965) | lac operon in E. coli |
| Gene fine structure | Benzer (phage T4) | Recon < Muton < Cistron |
| DNA model | Watson & Crick (1953; Nobel 1962) | Double helix; 3.4 Å/bp; 10 bp/turn |
| Chargaff’s Rules | A=T, G=C | Base pair ratio is species-specific |
| Genetic code | Triplet, degenerate, universal | 64 codons for 20 amino acids |
| Start codon | AUG (methionine) | Initiates translation |
| Stop codons | UAA, UAG, UGA | Ochre, Amber, Opal |
| Degeneracy | Multiple codons per amino acid | Wobble position provides mutation buffer |
Summary Cheat Sheet
| Concept / Topic | Key Details |
|---|---|
| Nucleic acids first isolated by | Miescher (1868) — “nuclein” from WBC pus |
| DNA proved as genetic material | Avery, MacLeod & McCarty (1944) |
| In vitro DNA synthesis | A. Kornberg (Nobel 1959); discovered DNA polymerase |
| In vitro RNA synthesis | S. Ochoa (Nobel 1959); polynucleotide phosphorylase |
| Artificial gene synthesis | Khorana & Agrawal — alanine tRNA gene from yeast |
| One gene–one enzyme | Beadle & Tatum (1943) on Neurospora crassa |
| Modern refinement | One gene → one polypeptide |
| Operon concept | Jacob & Monod (Nobel 1965); lac operon in E. coli |
| Benzer’s units (size order) | Recon (1–2 bp) < Muton (1 bp) < Cistron (hundreds bp) |
| Recon | Smallest unit of recombination |
| Muton | Smallest unit of mutation |
| Cistron | Functional unit = gene; codes for one polypeptide |
| DNA double helix | Watson & Crick (1953); Nobel Prize 1962 |
| X-ray data by | Wilkins & Rosalind Franklin (Photo 51) |
| A–T = 2 H-bonds | G–C = 3 H-bonds (more stable) |
| DNA dimensions | 3.4 Å/bp, 10 bp/turn, 34 Å/turn, 20 Å diameter |
| Chargaff’s Rules | A=T, G=C; (A+T)/(G+C) = species-specific ratio |
| RNA vs DNA | RNA: ribose, single-stranded, uracil replaces thymine |
| RdRp | RNA-dependent RNA polymerase; replicates viral RNA genomes |
| Genetic code deciphered by | Holley, Khorana, Nirenberg (Nobel 1968) |
| Triplet code | 4³ = 64 codons for 20 amino acids |
| Code properties | Triplet, Degenerate, Non-overlapping, Comma-less, Universal |
| Start codon | AUG (methionine) |
| Stop codons | UAA (ochre), UAG (amber), UGA (opal) |
Knowledge Check
Take a dynamically generated quiz based on the material you just read to test your understanding and get personalized feedback.
Lesson Doubts
Ask questions, get expert answers