☄️Enzymes — Meaning, Characteristics, Types, and Mode of Action
Enzyme discovery, characteristics, Michaelis-Menten constant, active and allosteric sites, classification by structure and function, and enzyme nomenclature with exam-focused tables
From Field to Lab — Enzymes in Everyday Agriculture
In the previous chapter on respiration, every single step — from glycolysis to the Krebs cycle to ETS — was catalysed by specific enzymes. Without enzymes, these reactions would take years instead of milliseconds. Now we study the catalysts themselves.
When a farmer adds urea to a rice field, the soil enzyme urease breaks it down into ammonium — a form the plant can absorb. When barley seeds are malted for brewing, the enzyme amylase converts stored starch into fermentable sugars. When milk curdles to form curd, Lactobacillus bacteria produce lactic acid through fermentation, which lowers the pH and causes milk proteins to coagulate. Every biological process in agriculture — from seed germination to soil nutrient cycling — is driven by enzymes.
This lesson covers:
- Discovery milestones — Buchner, Sumner, Kuhne, and ribozymes
- What enzymes are — biocatalysts that lower activation energy
- Characteristics — specificity, sensitivity, colloidal nature
- Catalytic efficiency — turn over number and Michaelis-Menten constant (Km)
- Active and allosteric sites — substrate binding vs regulation
- Types of enzymes — simple, conjugated, isoenzymes, constitutive, induced
- IUB classification — the 6 classes of enzymes
Understanding enzyme characteristics, types, and naming conventions is essential for biochemistry questions in competitive exams.
Discovery of Enzymes
The understanding that biological reactions are driven by specific protein catalysts developed over more than a century. These milestones are frequently tested in exams.
| Milestone | Scientist | Detail |
|---|---|---|
| First enzyme isolated | Buchner (1897) | Isolated Zymase from yeast; proved fermentation occurs in cell-free extracts (Nobel Prize 1907) |
| First purified and crystallised enzyme | J.B. Sumner (1926) | Crystallised Urease from Canavalia (Jack bean / Lobia); proved enzymes are proteins |
| Term “enzyme” coined | W. Kuhne (1878) | Greek: En = In, Zyme = Living (coined while studying fermentation) |
| First ribozyme discovered | T. Cech | L19 RNAase from Tetrahymena (protozoan); Altman discovered RNAase P in prokaryotes (Nobel Prize 1989) |
IMPORTANT
Ribozymes are RNA molecules that can catalyse chemical reactions — they challenged the assumption that only proteins can be enzymes. This discovery was key to understanding the origin of life (RNA world hypothesis).
What Are Enzymes?
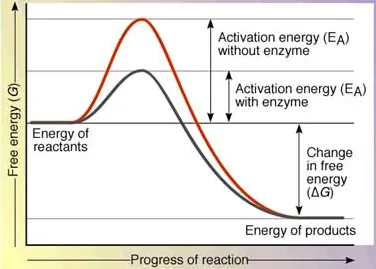
Enzymes are biocatalysts made up of proteins (except ribozymes) that increase the rate of chemical reactions by lowering the activation energy — the minimum energy needed for a reaction to proceed.
- Enzymes do not change the equilibrium of a reaction — they only speed up the rate at which equilibrium is reached
- Enzymes accelerate reactions by factors of millions to billions without being consumed
Characteristics of Enzymes
Enzymes share a set of properties that distinguish them from inorganic catalysts. These characteristics explain why biological reactions are so precisely controlled and why environmental stress (heat, pH change) can shut down metabolism.
| Characteristic | Detail | Agricultural Relevance |
|---|---|---|
| Chemical nature | Almost all are proteins | Protein-deficient plants have poor enzyme activity |
| Physical nature | Colloidal substances | Large surface area for substrate interaction |
| Sensitivity | Very sensitive to pH and temperature | Optimum temperature: 20–35°C |
| Synthesis | On ribosomes under gene control | Mutations can produce defective enzymes |
| Structure | Tertiary and globular proteins; isoenzymes are quaternary | 3D folding creates the active site |
| Quantity needed | Very minute amounts | High catalytic efficiency |
| Specificity | Very specific to substrate | Each enzyme fits only its substrate (“lock and key”) |
Catalytic Efficiency — Turn Over Number and Km
Two quantitative measures define how well an enzyme performs. The Turn Over Number measures raw speed, while the Michaelis-Menten constant (Km) measures substrate affinity. Both are commonly tested.
Turn Over Number
The turn over number (also called kcat) is the number of substrate molecules converted to products per unit time by a single enzyme molecule. Carbonic anhydrase holds the record — it processes 36 million molecules per minute, making it one of the fastest enzymes known.
| Enzyme | Turn Over Number (per minute) |
|---|---|
| Carbonic Anhydrase | 36,000,000 (highest known) |
| Catalase | 5,000,000 |
| Flavoprotein | 50 |
| Lysozyme | 30 |
Michaelis-Menten Constant (Km)
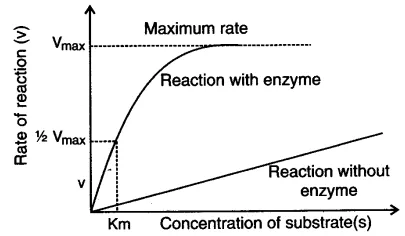
Km = substrate concentration at which the reaction rate is half of Vmax.
- A low Km = high affinity (enzyme reaches half-max at low substrate)
- A high Km = low affinity (needs more substrate)
- Given by Michaelis & Menten
TIP
Exam shortcut: Low Km = high affinity = efficient enzyme. Think of it as: the enzyme is so good at its job that it needs very little substrate to work at half-speed.
Active Site and Allosteric Site
Enzymes have two functionally important regions. The active site is where catalysis happens, while the allosteric site is where regulatory molecules bind to increase or decrease enzyme activity. This distinction is critical for understanding metabolic regulation (e.g., feedback inhibition in biosynthetic pathways).
| Feature | Active Site | Allosteric Site |
|---|---|---|
| What binds | Substrate | Modulators (activators or inhibitors) |
| Specificity | Determined by specific amino acid sequence | Different from active site (“allo” = other, “stereos” = shape) |
| Effect of binding | Catalysis of reaction | Modifies enzyme activity (regulation) |
| Examples | Substrate binding to hexokinase | Feedback inhibitors binding to PFK |
Types of Enzymes
Enzymes are classified in multiple ways — by their structural composition (simple vs conjugated), by when they are produced (constitutive vs induced), and by the type of reaction they catalyse (IUB classification). Each classification system is tested independently in exams.
A. By Structure
| Type | Composition | Examples |
|---|---|---|
| Simple enzymes | Wholly made of protein | Trypsin, pepsin, amylase, urease |
| Conjugated enzymes | Protein (apoenzyme) + non-protein (cofactor) = holoenzyme | Cytochrome oxidase, dehydrogenases |
| Isoenzymes | Different molecular structure but same function | Lactic Acid Dehydrogenase (LDH) |
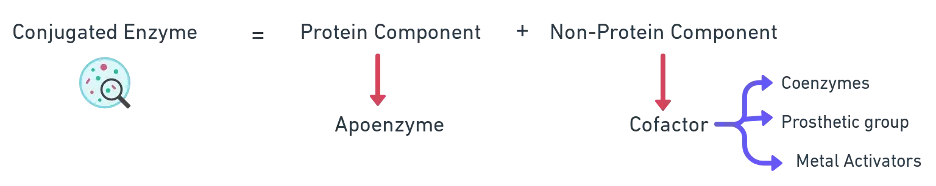
Cofactors come in three forms, distinguished by how tightly they bind and whether they are organic or inorganic:
| Co-enzyme | Prosthetic group | Metal Activators |
|---|---|---|
| Loosely attached organic molecule that can be separated by dialysis. Examples: Vitamins, AMP, ADP, NAD, NADP, FAD, Lipoic acid. | Tightly attached organic (sometimes inorganic) group that cannot be easily separated from the enzyme. Examples: Haem group in catalase and peroxidase enzymes. | Loosely attached inorganic or metallic ions that activate the enzyme. Examples: Fe²⁺, Zn²⁺, Mg²⁺ |
Conjugated enzyme equation:
Holoenzyme = Apoenzyme (protein) + Cofactor (non-protein)
| Cofactor Type | Nature | Examples |
|---|---|---|
| Prosthetic group | Tightly bound non-protein organic molecule | FAD, FMN, haem |
| Co-enzyme | Loosely bound non-protein organic molecule | NAD⁺, NADP⁺, CoA |
| Metal ion activator | Inorganic ion | Zn²⁺, Mg²⁺, Fe²⁺, Mn²⁺ |
TIP
Mnemonic — “HAC”: Holoenzyme = Apoenzyme + Cofactor. The complete enzyme needs both its protein part and its helper.
B. By Presence
| Type | Present When | Example |
|---|---|---|
| Constitutive enzymes | Always present in cell regardless of need | Glycolytic enzymes |
| Induced enzymes | Produced only when substrate is present | Lactase in bacteria (induced by lactose) |
Enzyme Nomenclature — IUB Classification
The International Union of Biochemistry (IUB) provides a systematic classification that assigns every enzyme a 4-digit EC number and places it in one of 6 classes based on the type of reaction it catalyses. This system replaces trivial names (like “pepsin” or “trypsin”) with systematic names, though trivial names are still widely used in exams.
| Class | Reaction Type | Example | Memory Aid |
|---|---|---|---|
| 1. Oxidoreductases | Oxidation-reduction (electron transfer) | Dehydrogenase | Redox |
| 2. Transferases | Transfer of functional groups | Transaminase, Kinase | Transfer |
| 3. Hydrolases | Hydrolysis (breaking bonds with water) | Lipase, Amylase, Protease | Hydro = water |
| 4. Lyases | Breaking bonds without water or oxidation | Decarboxylase, Aldolase | Lyse = break |
| 5. Isomerases | Isomer inter-conversion | Phosphoglucose isomerase | Iso = same |
| 6. Ligases | Joining two molecules using ATP | DNA ligase, Carboxylase | Liga = bind |
TIP
Mnemonic — “Over The Hill Lies Incredible Land”: Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases. These 6 classes cover all known enzymes.
Summary Table — Key Facts at a Glance
| Fact | Answer |
|---|---|
| First enzyme discovered | Zymase (from yeast) by Buchner |
| First crystallised enzyme | Urease by J.B. Sumner (from Jack bean) |
| Term “enzyme” coined by | W. Kuhne (1878) |
| All enzymes are | Proteins (except ribozymes) |
| Enzymes are | Colloidal in nature |
| Optimum temperature | 20–35°C |
| Highest turn over number | Carbonic Anhydrase (36 million/min) |
| Low Km means | High affinity for substrate |
| Holoenzyme = | Apoenzyme + Cofactor |
| Ribozymes are | RNA catalysts (Cech, Altman — Nobel 1989) |
| 6 enzyme classes | Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases |
| Isoenzymes | Different structure, same function |
| Active site | Where substrate binds |
| Allosteric site | Where modulators bind (regulation) |
Summary Cheat Sheet
| Fact | Answer |
|---|---|
| Term “enzyme” coined by | W. Kuhne (1878) — Greek: En = In, Zyme = Living |
| First enzyme isolated | Zymase by Buchner (1897) from yeast |
| First crystallised enzyme | Urease by J.B. Sumner (1926) from Jack bean (Canavalia) |
| Ribozyme discoverers | T. Cech & Altman (Nobel Prize 1989) |
| Ribozymes are made of | RNA (not protein) |
| Enzymes lower | Activation energy (do NOT change equilibrium) |
| Chemical nature of enzymes | Proteins (except ribozymes) |
| Physical nature of enzymes | Colloidal substances |
| Enzyme structure type | Tertiary and globular proteins |
| Optimum temperature for enzymes | 20–35°C |
| Highest turn over number | Carbonic Anhydrase — 36 million/min |
| Km definition | Substrate concentration at half of Vmax |
| Low Km means | High affinity for substrate |
| Km given by | Michaelis & Menten |
| Holoenzyme formula | Apoenzyme (protein) + Cofactor (non-protein) |
| Prosthetic group binding | Tightly bound organic molecule (e.g., FAD, haem) |
| Co-enzyme binding | Loosely bound organic molecule (e.g., NAD⁺, CoA) |
| Isoenzymes | Different structure, same function (e.g., LDH) |
| Constitutive enzymes | Always present regardless of substrate |
| Induced enzymes | Produced only when substrate is present |
| IUB 6 enzyme classes | Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases |
| Ligases require | ATP to join two molecules |
| Active site binds | Substrate (catalysis) |
| Allosteric site binds | Modulators (regulation) |
TIP
Next: Lesson 04-02 covers the factors affecting enzyme activity — temperature, pH, substrate concentration, inhibitors (competitive and non-competitive), and enzyme regulation mechanisms.
Pro Content Locked
Upgrade to Pro to access this lesson and all other premium content.
₹2388 billed yearly
- All Agriculture & Banking Courses
- AI Lesson Questions (100/day)
- AI Doubt Solver (50/day)
- Glows & Grows Feedback (30/day)
- AI Section Quiz (20/day)
- 22-Language Translation (30/day)
- Recall Questions (20/day)
- AI Quiz (15/day)
- AI Quiz Paper Analysis
- AI Step-by-Step Explanations
- Spaced Repetition Recall (FSRS)
- AI Tutor
- Immersive Text Questions
- Audio Lessons — Hindi & English
- Mock Tests & Previous Year Papers
- Summary & Mind Maps
- XP, Levels, Leaderboard & Badges
- Generate New Classrooms
- Voice AI Teacher (AgriDots Live)
- AI Revision Assistant
- Knowledge Gap Analysis
- Interactive Revision (LangGraph)
🔒 Secure via Razorpay · Cancel anytime · No hidden fees
From Field to Lab — Enzymes in Everyday Agriculture
In the previous chapter on respiration, every single step — from glycolysis to the Krebs cycle to ETS — was catalysed by specific enzymes. Without enzymes, these reactions would take years instead of milliseconds. Now we study the catalysts themselves.
When a farmer adds urea to a rice field, the soil enzyme urease breaks it down into ammonium — a form the plant can absorb. When barley seeds are malted for brewing, the enzyme amylase converts stored starch into fermentable sugars. When milk curdles to form curd, Lactobacillus bacteria produce lactic acid through fermentation, which lowers the pH and causes milk proteins to coagulate. Every biological process in agriculture — from seed germination to soil nutrient cycling — is driven by enzymes.
This lesson covers:
- Discovery milestones — Buchner, Sumner, Kuhne, and ribozymes
- What enzymes are — biocatalysts that lower activation energy
- Characteristics — specificity, sensitivity, colloidal nature
- Catalytic efficiency — turn over number and Michaelis-Menten constant (Km)
- Active and allosteric sites — substrate binding vs regulation
- Types of enzymes — simple, conjugated, isoenzymes, constitutive, induced
- IUB classification — the 6 classes of enzymes
Understanding enzyme characteristics, types, and naming conventions is essential for biochemistry questions in competitive exams.
Discovery of Enzymes
The understanding that biological reactions are driven by specific protein catalysts developed over more than a century. These milestones are frequently tested in exams.
| Milestone | Scientist | Detail |
|---|---|---|
| First enzyme isolated | Buchner (1897) | Isolated Zymase from yeast; proved fermentation occurs in cell-free extracts (Nobel Prize 1907) |
| First purified and crystallised enzyme | J.B. Sumner (1926) | Crystallised Urease from Canavalia (Jack bean / Lobia); proved enzymes are proteins |
| Term “enzyme” coined | W. Kuhne (1878) | Greek: En = In, Zyme = Living (coined while studying fermentation) |
| First ribozyme discovered | T. Cech | L19 RNAase from Tetrahymena (protozoan); Altman discovered RNAase P in prokaryotes (Nobel Prize 1989) |
IMPORTANT
Ribozymes are RNA molecules that can catalyse chemical reactions — they challenged the assumption that only proteins can be enzymes. This discovery was key to understanding the origin of life (RNA world hypothesis).
What Are Enzymes?
Enzymes are biocatalysts made up of proteins (except ribozymes) that increase the rate of chemical reactions by lowering the activation energy — the minimum energy needed for a reaction to proceed.
- Enzymes do not change the equilibrium of a reaction — they only speed up the rate at which equilibrium is reached
- Enzymes accelerate reactions by factors of millions to billions without being consumed

Characteristics of Enzymes
Enzymes share a set of properties that distinguish them from inorganic catalysts. These characteristics explain why biological reactions are so precisely controlled and why environmental stress (heat, pH change) can shut down metabolism.
| Characteristic | Detail | Agricultural Relevance |
|---|---|---|
| Chemical nature | Almost all are proteins | Protein-deficient plants have poor enzyme activity |
| Physical nature | Colloidal substances | Large surface area for substrate interaction |
| Sensitivity | Very sensitive to pH and temperature | Optimum temperature: 20–35°C |
| Synthesis | On ribosomes under gene control | Mutations can produce defective enzymes |
| Structure | Tertiary and globular proteins; isoenzymes are quaternary | 3D folding creates the active site |
| Quantity needed | Very minute amounts | High catalytic efficiency |
| Specificity | Very specific to substrate | Each enzyme fits only its substrate (“lock and key”) |
Catalytic Efficiency — Turn Over Number and Km
Two quantitative measures define how well an enzyme performs. The Turn Over Number measures raw speed, while the Michaelis-Menten constant (Km) measures substrate affinity. Both are commonly tested.
Turn Over Number
The turn over number (also called kcat) is the number of substrate molecules converted to products per unit time by a single enzyme molecule. Carbonic anhydrase holds the record — it processes 36 million molecules per minute, making it one of the fastest enzymes known.
| Enzyme | Turn Over Number (per minute) |
|---|---|
| Carbonic Anhydrase | 36,000,000 (highest known) |
| Catalase | 5,000,000 |
| Flavoprotein | 50 |
| Lysozyme | 30 |
Michaelis-Menten Constant (Km)
Km = substrate concentration at which the reaction rate is half of Vmax.
- A low Km = high affinity (enzyme reaches half-max at low substrate)
- A high Km = low affinity (needs more substrate)
- Given by Michaelis & Menten

TIP
Exam shortcut: Low Km = high affinity = efficient enzyme. Think of it as: the enzyme is so good at its job that it needs very little substrate to work at half-speed.
Active Site and Allosteric Site
Enzymes have two functionally important regions. The active site is where catalysis happens, while the allosteric site is where regulatory molecules bind to increase or decrease enzyme activity. This distinction is critical for understanding metabolic regulation (e.g., feedback inhibition in biosynthetic pathways).
| Feature | Active Site | Allosteric Site |
|---|---|---|
| What binds | Substrate | Modulators (activators or inhibitors) |
| Specificity | Determined by specific amino acid sequence | Different from active site (“allo” = other, “stereos” = shape) |
| Effect of binding | Catalysis of reaction | Modifies enzyme activity (regulation) |
| Examples | Substrate binding to hexokinase | Feedback inhibitors binding to PFK |
Types of Enzymes
Enzymes are classified in multiple ways — by their structural composition (simple vs conjugated), by when they are produced (constitutive vs induced), and by the type of reaction they catalyse (IUB classification). Each classification system is tested independently in exams.
A. By Structure
| Type | Composition | Examples |
|---|---|---|
| Simple enzymes | Wholly made of protein | Trypsin, pepsin, amylase, urease |
| Conjugated enzymes | Protein (apoenzyme) + non-protein (cofactor) = holoenzyme | Cytochrome oxidase, dehydrogenases |
| Isoenzymes | Different molecular structure but same function | Lactic Acid Dehydrogenase (LDH) |

Cofactors come in three forms, distinguished by how tightly they bind and whether they are organic or inorganic:
| Co-enzyme | Prosthetic group | Metal Activators |
|---|---|---|
| Loosely attached organic molecule that can be separated by dialysis. Examples: Vitamins, AMP, ADP, NAD, NADP, FAD, Lipoic acid. | Tightly attached organic (sometimes inorganic) group that cannot be easily separated from the enzyme. Examples: Haem group in catalase and peroxidase enzymes. | Loosely attached inorganic or metallic ions that activate the enzyme. Examples: Fe²⁺, Zn²⁺, Mg²⁺ |
Conjugated enzyme equation:
Holoenzyme = Apoenzyme (protein) + Cofactor (non-protein)
| Cofactor Type | Nature | Examples |
|---|---|---|
| Prosthetic group | Tightly bound non-protein organic molecule | FAD, FMN, haem |
| Co-enzyme | Loosely bound non-protein organic molecule | NAD⁺, NADP⁺, CoA |
| Metal ion activator | Inorganic ion | Zn²⁺, Mg²⁺, Fe²⁺, Mn²⁺ |
TIP
Mnemonic — “HAC”: Holoenzyme = Apoenzyme + Cofactor. The complete enzyme needs both its protein part and its helper.
B. By Presence
| Type | Present When | Example |
|---|---|---|
| Constitutive enzymes | Always present in cell regardless of need | Glycolytic enzymes |
| Induced enzymes | Produced only when substrate is present | Lactase in bacteria (induced by lactose) |
Enzyme Nomenclature — IUB Classification
The International Union of Biochemistry (IUB) provides a systematic classification that assigns every enzyme a 4-digit EC number and places it in one of 6 classes based on the type of reaction it catalyses. This system replaces trivial names (like “pepsin” or “trypsin”) with systematic names, though trivial names are still widely used in exams.
| Class | Reaction Type | Example | Memory Aid |
|---|---|---|---|
| 1. Oxidoreductases | Oxidation-reduction (electron transfer) | Dehydrogenase | Redox |
| 2. Transferases | Transfer of functional groups | Transaminase, Kinase | Transfer |
| 3. Hydrolases | Hydrolysis (breaking bonds with water) | Lipase, Amylase, Protease | Hydro = water |
| 4. Lyases | Breaking bonds without water or oxidation | Decarboxylase, Aldolase | Lyse = break |
| 5. Isomerases | Isomer inter-conversion | Phosphoglucose isomerase | Iso = same |
| 6. Ligases | Joining two molecules using ATP | DNA ligase, Carboxylase | Liga = bind |
TIP
Mnemonic — “Over The Hill Lies Incredible Land”: Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases. These 6 classes cover all known enzymes.
Summary Table — Key Facts at a Glance
| Fact | Answer |
|---|---|
| First enzyme discovered | Zymase (from yeast) by Buchner |
| First crystallised enzyme | Urease by J.B. Sumner (from Jack bean) |
| Term “enzyme” coined by | W. Kuhne (1878) |
| All enzymes are | Proteins (except ribozymes) |
| Enzymes are | Colloidal in nature |
| Optimum temperature | 20–35°C |
| Highest turn over number | Carbonic Anhydrase (36 million/min) |
| Low Km means | High affinity for substrate |
| Holoenzyme = | Apoenzyme + Cofactor |
| Ribozymes are | RNA catalysts (Cech, Altman — Nobel 1989) |
| 6 enzyme classes | Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases |
| Isoenzymes | Different structure, same function |
| Active site | Where substrate binds |
| Allosteric site | Where modulators bind (regulation) |
Summary Cheat Sheet
| Fact | Answer |
|---|---|
| Term “enzyme” coined by | W. Kuhne (1878) — Greek: En = In, Zyme = Living |
| First enzyme isolated | Zymase by Buchner (1897) from yeast |
| First crystallised enzyme | Urease by J.B. Sumner (1926) from Jack bean (Canavalia) |
| Ribozyme discoverers | T. Cech & Altman (Nobel Prize 1989) |
| Ribozymes are made of | RNA (not protein) |
| Enzymes lower | Activation energy (do NOT change equilibrium) |
| Chemical nature of enzymes | Proteins (except ribozymes) |
| Physical nature of enzymes | Colloidal substances |
| Enzyme structure type | Tertiary and globular proteins |
| Optimum temperature for enzymes | 20–35°C |
| Highest turn over number | Carbonic Anhydrase — 36 million/min |
| Km definition | Substrate concentration at half of Vmax |
| Low Km means | High affinity for substrate |
| Km given by | Michaelis & Menten |
| Holoenzyme formula | Apoenzyme (protein) + Cofactor (non-protein) |
| Prosthetic group binding | Tightly bound organic molecule (e.g., FAD, haem) |
| Co-enzyme binding | Loosely bound organic molecule (e.g., NAD⁺, CoA) |
| Isoenzymes | Different structure, same function (e.g., LDH) |
| Constitutive enzymes | Always present regardless of substrate |
| Induced enzymes | Produced only when substrate is present |
| IUB 6 enzyme classes | Oxidoreductases, Transferases, Hydrolases, Lyases, Isomerases, Ligases |
| Ligases require | ATP to join two molecules |
| Active site binds | Substrate (catalysis) |
| Allosteric site binds | Modulators (regulation) |
TIP
Next: Lesson 04-02 covers the factors affecting enzyme activity — temperature, pH, substrate concentration, inhibitors (competitive and non-competitive), and enzyme regulation mechanisms.
Knowledge Check
Take a dynamically generated quiz based on the material you just read to test your understanding and get personalized feedback.
Lesson Doubts
Ask questions, get expert answers